Drug development has long been a process of trial-and-error to test biological hypotheses against clinical reality. Despite advancements in science and technology, the timeline from target ID to a clinical candidate still often takes more than five years, and nearly 90% of drugs that enter clinical trials fail. With treatment landscapes evolving and modalities becoming increasingly complex, it is no surprise that R&D costs per approved therapy continue to double every nine years. The limiting factor in drug development has never been a shortage of hypotheses, but rather a shortage of the resources to evaluate them effectively and efficiently.
Machine learning in drug design holds the promise to change that math by accelerating iteration and improving the odds of success. Between 2012 and 2022, approximately 200 companies that leveraged AI for drug discovery
raised a collective $18B. We are now seeing the results of these efforts play out in the clinic.
In June 2025, Insilico Medicine
published positive Phase IIa results in
Nature Medicine for its first-in-class small molecule
TNIK inhibitor rentosertib in idiopathic pulmonary fibrosis. This made it the first drug to generate clinical proof-of-concept for which both the target was discovered and the molecule was designed entirely using generative AI. In this example, AI played a critical role in changing “the math” by leveraging a generative chemistry platform for molecule design and optimization. The team nominated a preclinical candidate after screening only 78 molecules, rather than the thousands typically required, and did so
in 18 months at less than 10% of the average cost per approved drug.
With a favorable investment-reward profile, it is no surprise that many companies, including large pharma, have made concerted efforts to incorporate AI platforms into the R&D process to accelerate drug discovery. In early 2026, GSK and Eli Lilly announced deals with
NOETIK and
Chai Discovery for access to their oncology and drug design foundation models, with GSK committing $50M upfront to NOETIK and Lilly paying a mid-eight-figure annual access fee to Chai for biologics design.
insitro, which integrates large-scale human cell data generation with machine learning, recently saw BMS nominate two additional ALS targets from their collaboration, validation of the value of the full-stack approach in pairing proprietary data generation with drug development.
Isomorphic Labs, the Google DeepMind spinout behind AlphaFold, has pursued deep partnerships with Lilly, Novartis, and J&J with potential value exceeding $3B, while advancing its own internal oncology pipeline toward first-in-human trials. Its newly released IsoDDE model more than doubles AlphaFold 3's accuracy on the hardest generalization benchmarks, making it one of the most closely watched companies in AI-driven drug design. And it's not just pharma showing interest: in early April 2026, Anthropic acquired
Coefficient Bio, an eight-month-old startup built by former Evozyne/Genentech/Prescient Design computational biologists, for $400M in stock, signaling that frontier AI labs are now making direct bets on drug discovery.
Though computational chemistry tools first emerged in the 1980s, the modern era of AI for biotech effectively began with the rise of deep learning in the 2010s, when it became clear that neural networks could learn meaningful representations of molecular structure from data. The watershed moment occurred when DeepMind's AlphaFold2 and the Baker Lab's RoseTTAFold solved the problem of predicting a protein’s 3D structure from its amino acid sequence alone. From there, the number of biological AI models grew exponentially. By 2024, over 350 biological AI models were published, including AlphaFold3, ESM3, Boltz-1, BindCraft, Evo, scGPT, and H-Optimus-0, highlighting AI’s ability to perform tasks across generative protein design, genomics and perturbation modeling, and pathology image analysis.
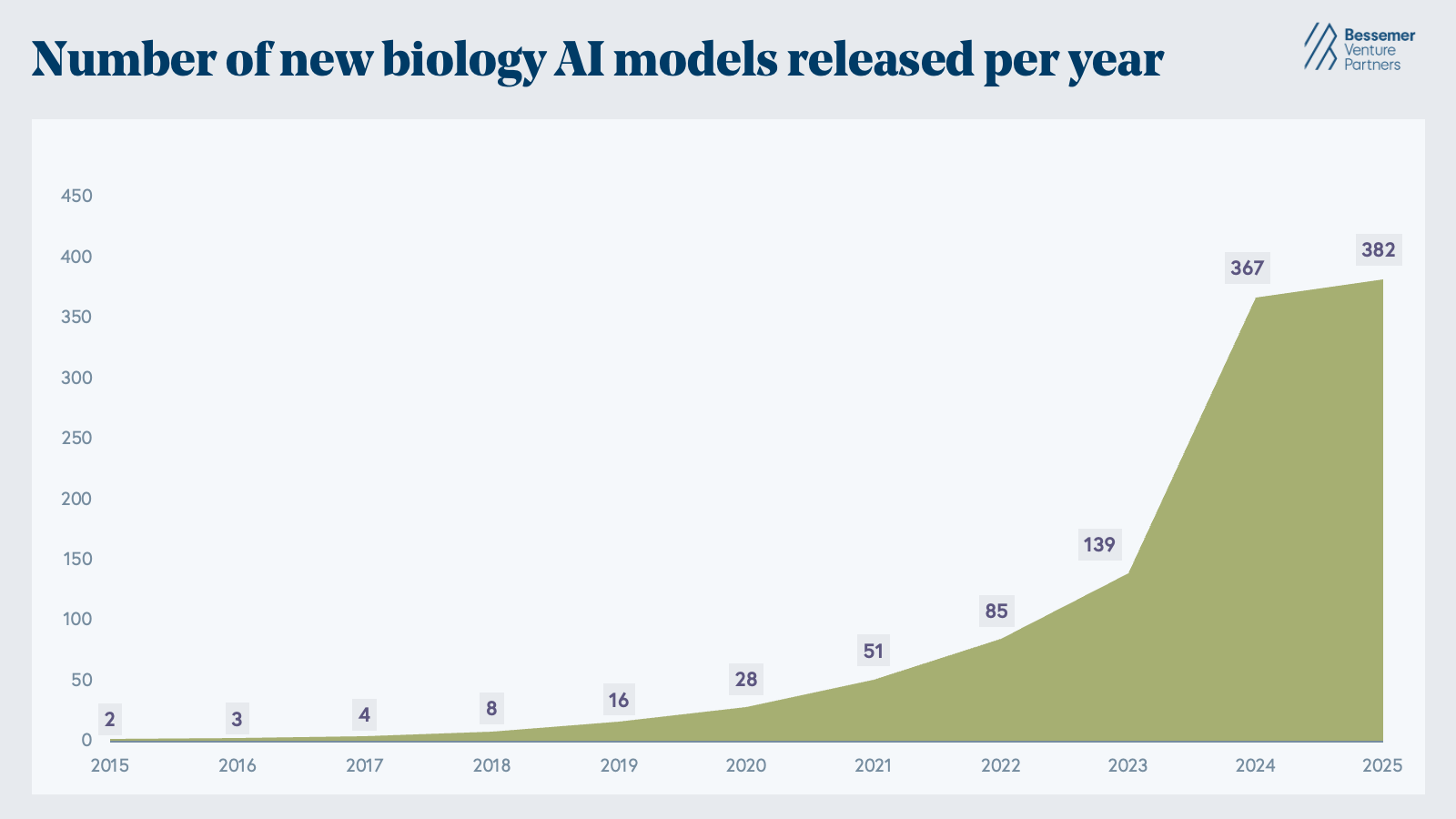
The Cambrian explosion of AI models for biology has already happened. Between 2015 and 2025, the number of new biology AI models released each year has exponentially increased from sub-ten to 380+ and counting. (Note that this is based purely on the dataset from Epoch AI and may not be complete.)
Most recently, new models such as JAM-2, BoltzGen, Latent-X2, Chai-2, and IsoDDE have continued to bring us closer to designing drug-like biologics straight from the computer. Momentum on the zero-shot design task has never been stronger. Following a surge in new AI models for biology, the field now has an armamentarium of tools spanning the drug development continuum, from structural modeling to molecule design and drug optimization.
Three principles of biology-native data infrastructure
In an increasingly crowded landscape, we believe the AI-driven biotechs that persist and scale over time will be those built on three core principles, which together we define as the principles of biology-native data infrastructure:
- Curating scalable, multi-modal datasets informed by the biological challenges associated with a drug’s mechanism of action.
- Incorporating the newest agentic AI frameworks across entire R&D workflows.
- Adopting lab automation to power rapid, closed experimental feedback loops.
Companies that enable or embody these principles will be the ones to truly accelerate drug design timelines, reduce clinical trial failure risk, and deliver on the promise that AI in biology holds.
Below we explain why these principles are critical to the drug development industry and highlight the emerging categories and companies that are putting these principles into practice.
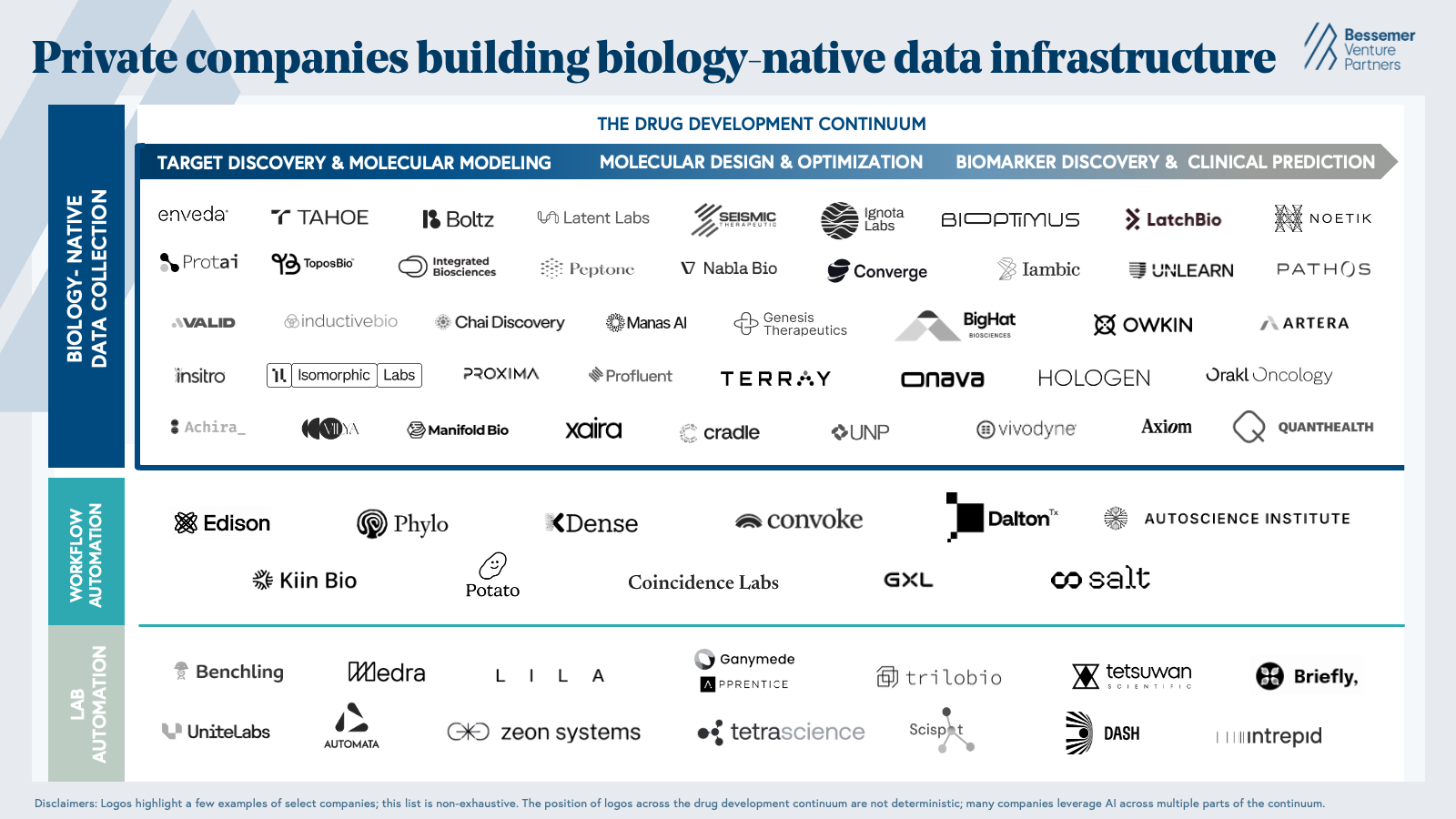
Market map
Our market map highlights private life science companies that are leveraging AI to create and analyze biological datasets that address challenges along the drug development continuum, accelerate R&D workflows end-to-end, and automate the physical work of conducting wet lab experiments.
1. Biology-native data at scale
Much of the data that makes current AI biology models possible was slowly assembled over decades of publicly-funded science. The Protein Data Bank’s (PDB) 200K+ protein structures were experimentally determined by techniques like X-ray crystallography and NMR spectroscopy. Similarly, the Human Genome Project’s map of human genes and DNA was the result of sequencing efforts across global research institutions, and ChEMBL’s bioactivity database on millions of small molecules was accumulated through years of manual patent and literature data extraction. The impact of these databases is remarkable—for example, structural data from the PDB contributed to the development of 100% of the protein-targeted small-molecule cancer drugs approved by the FDA between 2019 and 2023.
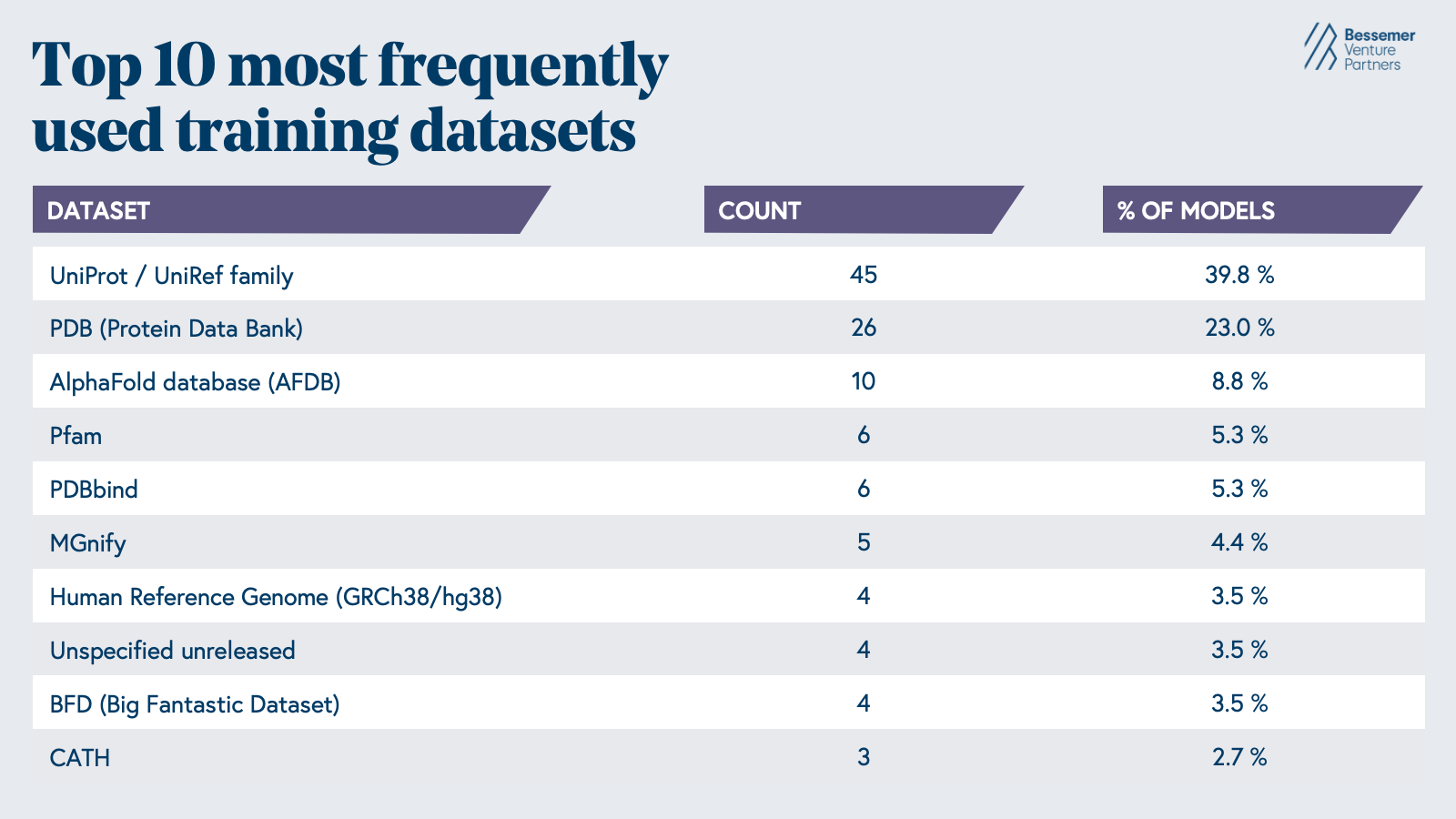
The AI biology models developed in the last few decades reflect the data that is readily available, with nearly 63% of models being trained on protein sequences and structures from the Uniprot database and the PDB (
Epoch AI).
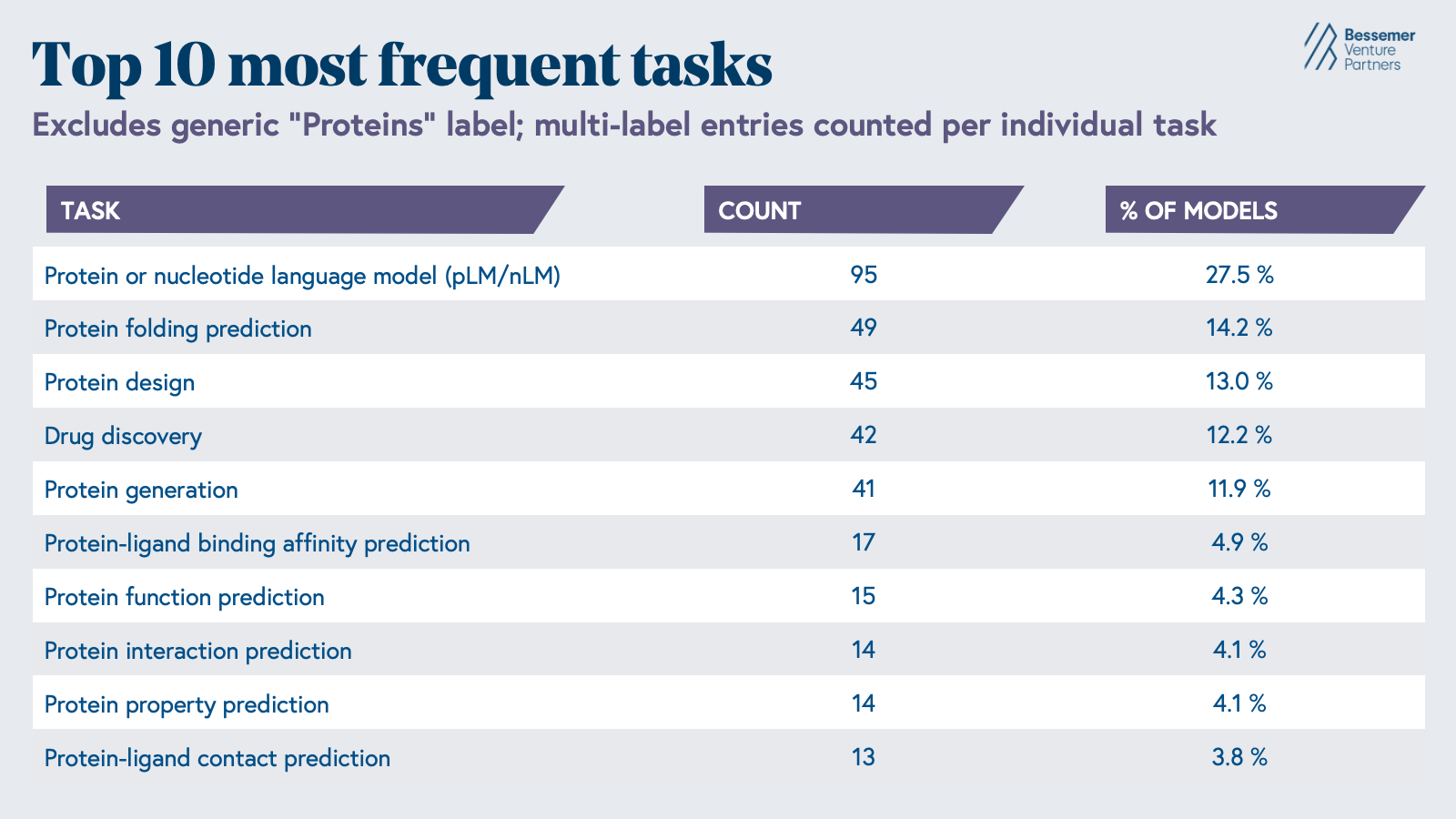
The most common tasks these models are used for are the contextual understanding of protein or nucleotide sequences, protein folding prediction, or protein design. However, significant gaps in our understanding of early-stage drug discovery biology persist, driven by the sheer complexity of biological systems and the limitations of the tools we have to study them.
Despite its scale, the PDB is heavily biased toward proteins that are stable, soluble, and amenable to crystallization. Though membrane proteins, intrinsically disordered proteins, and transient protein complexes are some of the most compelling drug targets in oncology and neurodegeneration, they often defy these criteria and therefore remain dramatically underrepresented. Additionally, the structures the PDB captures are static snapshots, freezing proteins in a single conformation rather than the dynamic ensemble of shapes they adopt in a living cell. Yet it is often these alternative conformations that are most therapeutically relevant, as seen with allosteric binding sites that only become accessible upon ligand binding.
Although bringing a new drug to market begins with protein structure and design tasks, early-stage drug discovery accounts for only a fraction of the time and cost of the drug development process.
More than two-thirds of drug development time and resources are allocated to the steps after early drug discovery, including the ADME (pharmacokinetic properties related to “absorption, distribution, metabolism, and excretion”) and formulation optimization work done in preclinical studies, as well as the safety and efficacy studies run in clinical trials. To advance a drug from a hit to lead to a development candidate, far more is needed than confirmation that a molecule binds its target. The drug development process also requires an understanding of developability, immunogenicity, off-target effects, thermostability, solubility, and aggregation propensity, properties for which large, high-quality public datasets to supervise model learning currently don't exist.
While drug discovery is fundamentally a problem of understanding perturbations, there is no PDB-equivalent repository for understanding cell phenotypes in response to perturbations, or even proteomics data across disease states. Tying cellular to clinical data presents an even greater gap, as patient-level omics profiles linked to treatment outcomes and trial responses exist siloed in hospital systems and biopharma databases, making it nearly impossible to train models that could predict which patients will respond to a given therapy before they ever enroll in a trial. These are precisely the properties that determine whether a molecule can eventually become an approved drug, which means the most commercially important predictions are also the ones where the data infrastructure is weakest.
Much of the biological data available today was generated before the explosion of AI biology models, meaning it often lacks the traits that make it useful for machine learning. Annotations are often incomplete or unstandardized, and important context, such as cellular environment or lab equipment used, is rarely captured or encoded into datasets. In many cases, biological datasets simply don’t have the scale for models to draw statistically significant conclusions or make unbiased predictions. And even where scale exists, data tends to be siloed by modality—genomic, transcriptomic, pathology, and clinical outcome datasets frequently are collected and live in separate places, making it challenging to construct a data layer that allows AI to reason across the full picture of human biology.
| To truly unlock AI for drug development, we believe companies should be investing on two fronts: first in generating novel, multi-modal biological measurements that broaden our understanding of disease and second in building datasets with the scale, consistency, and contextualization needed to train models that generalize across diverse biological settings. |
We’re grateful to have backed several companies that exemplify this principle.
Peptone is combining atom-level biophysics with supercomputing to generate proprietary structural data on intrinsically disordered proteins, and
Inductive Bio is assembling one of the industry's largest and most diverse ADMET datasets to train its Beacon models,
which recently placed first among 370+ submissions in the OpenADMET-ExpansionRx endpoint prediction challenge.
Converge Bio is generating large-scale datasets to train and validate their own models to deploy with pharma and biotech customers for antibody design or sequence optimization, and
Seismic is taking a pipeline-first approach, using its IMPACT platform to parallelize the optimization of multiple drug-like properties of novel immunology biologics.
We’re also seeing progress downstream in the drug development continuum. For example,
NOETIK is assembling one of the most comprehensive datasets in oncology by pairing tumor multi-omics with longitudinal treatment outcomes, and
Prima Mente is building whole-genome epigenetic and multi-omic data models applied to brain disease. These data-rich disease-specific foundation models aim to enable novel target and biomarker discovery, more precise virtual cell simulation perturbation models, and improved clinical trial design.
2. Agentic AI across R&D workflows
While the cost of bringing a drug to market has increased, the cost of computing has decreased exponentially since the 1950s, consistent with Moore’s Law. Tasks along the drug development continuum that are computationally expensive today will be dramatically cheaper within a few years, and companies that build their tech stack to be rapidly adaptable to the evolving capabilities of AI will find themselves with an increasingly substantial structural advantage over those that view AI as a fixed investment.
The evolution of computational drug discovery workflows is a useful lens for what this adaptability looks like in practice. While building proprietary molecular modeling and simulation tools in-house may have been a differentiator a decade ago, the abundance of off-the-shelf in silico tools has changed this defensibility narrative. Structure predictors, ADMET models, and molecular dynamics simulators have greatly matured and are now widely accessible through both closed-source architectures and open-source repositories, often making it more time and resource-efficient to strategically mosaic tools across this ecosystem rather than build from scratch. The same logic applies as new foundation models emerge, new training techniques evolve, and new hardware enables greater compute efficiency.
| Companies should build their infrastructure from day one to be able to test, implement, and leverage the newest tools rather than be anchored to any single stack. Today, this modular infrastructure can look like a system that autonomously leverages and orchestrates the best tools for particular tasks, whether it’s literature review or running a bioinformatics pipeline. |
Cheaper compute has made long-context inference economically practical, enabling AI agents to synthesize over
1,000 papers and 40K lines of code in a single run. In combination with techniques that boost the accuracy and efficiency of AI, such as chain-of-thought reasoning and multi-agent frameworks, it has become increasingly realistic that AI can meaningfully compress the cost and time of the R&D lifecycle.
Agentic AI scientists could mine preprint servers, patent filings, and public biological databases to surface non-obvious connections, generate novel hypotheses, perform in silico data analysis, design wet lab experiments, and write reports, all while maintaining team-wide research context and a historical record of experiments that empowers scientists to make smarter and faster decisions.
Soon, it will be standard to adopt an AI operating system that spans the entire drug development process, leveraging AI’s ability to retain extensive context to unify analyses and results into a single research environment rather than leaving them siloed across disparate point solutions.
A growing wave of companies is building toward this vision, including both startups focused purely on the life sciences and frontier labs like Anthropic, which now offers
connectors to integrate Claude with platforms such as Benchling, PubMed, ChEMBL, ClinicalTrials.gov, and more.
K-Dense and
Edison Scientific are developing autonomous AI scientist platforms that can plan, execute, and iterate on complex, long-horizon research workflows end-to-end, from hypothesis generation to running computational experiments.
Phylo is taking a complementary approach with its Integrated Biology Environment, a unified workspace where scientists can seamlessly collaborate with AI agents across their datasets and analytical pipelines without switching between fragmented interfaces.
Companies like
Potato and
Convoke are building the operating systems for biopharma across early-stage drug discovery and downstream commercialization workflows, with Potato serving as the infrastructure to autonomously design and run experiments, and Convoke serving as a system of record and action to accelerate the regulatory and document-based workflows to bring drugs to market.
3. Closed loop lab automation
Even companies that utilize the most cutting-edge AI models run into the constraints of generating experimental data. Despite dramatic advances in structure prediction and molecular modeling, many in silico outputs, such as binding affinity predictions, still need to be validated in the wet lab before any downstream development decision can be made with confidence. Beyond that, in vivo efficacy is essentially unpredictable from first principles, with late-stage failures driven disproportionately by pharmacokinetic and toxicity properties that in silico models failed to flag. Given that experimental results are the ultimate source of biological ground truth, it’s crucial that these models continuously incorporate feedback from the wet lab to be anchored in accuracy.
Unfortunately, the experimental cycles that stand between a model's output and the data needed to update that model’s priors often take weeks to months. Wet lab experiments are slow, prone to failure, and dependent on skilled human labor, making them one of the biggest bottlenecks in shortening drug development timelines. The iterative design-test-make-analyze cycle that characterizes lead optimization can take up to three years itself and accounts for nearly a quarter of the total drug development timeline. These timelines are further extended by the reality that experimental validation is often outsourced to Contract Research Organizations (CROs), where coordination overhead, queue times, and data quality inconsistencies can add weeks or months to each iteration cycle. Bringing experimental capabilities in-house is increasingly necessary as it gives teams control over the context and caliber of data generation that makes closed-loop learning meaningful.
Though Hamilton robots for liquid handling and Chemspeed platforms for automated synthesis have existed in labs for decades, they’re optimized for the high throughput of specific point tasks rather than the automation and integration of entire experimental workflows. Most lab automation today still requires significant human intervention to transfer materials between instruments, troubleshoot failures, and interpret results before the next experimental step can begin, compressing individual tasks without compressing the end-to-end cycle.
In particular, automating robotic lab equipment has historically required dedicated automation engineers to configure the instruments and continuously write new scripts for different workflows. Natural language interfaces for robot control could effectively democratize automation capabilities, enabling scientists without any robotics or software engineering background to run, monitor, and iterate on experiments remotely and autonomously. Advancements in robotics and physical AI could further orchestrate the material and data transfer that is still being done by humans today. For example, vision-native systems can now autonomously read and interpret microscopy images of cells and feed structured data directly back into model pipelines without a scientist manually extracting and inputting the results.
Progress towards autonomous labs provides companies with significant leverage in both speed and operational spend. A model trained on five design-test-analyze cycles in the time a competitor completes one will compound its biological understanding far faster, and that compounding translates directly into better models, better molecules, and a structural advantage that is very difficult for those dependent on traditional CRO timelines to close. The companies building toward increased iteration speed through lab automation will also achieve greater data consistency, accuracy, and volume—reinforcing our first principle on the need for biology-native data at scale.
Companies across the lab automation landscape are pursuing this from different angles.
Medra is building an instrument-agnostic robotics platform where general-purpose robots interact with existing lab equipment through physical controls and software interfaces.
Automata takes a lab orchestration approach with its LINQ platform, providing modular hardware and software that connects disparate instruments into coordinated, end-to-end automated workflows.
Dash Bio is using robotics to become a faster and more automated CRO that offers the speed and consistency that in-house automation provides.
Lila Sciences represents one of the most vertically integrated approaches, building a fully automated lab for end-to-end drug discovery and development.
Life sciences will run on AI
We believe that companies building large biology-native datasets, AI-centric development stacks, and lab automation platforms that power rapid closed-loop experimentation will enable and define the next generation of life science companies.
We see this market organized across three interdependent layers. At the top sit companies generating data at the scale, modality, and fidelity AI requires to produce meaningful discoveries across the drug development continuum. Beneath them lie the physical and software infrastructure layers, including workflow and lab automation platforms, that compress timelines at every stage. Together, these three layers represent much of the value chain taking shape in AI-driven drug development, a core area of investment where we believe the next generation of life science companies will be built.
If you are building in any of these categories, or more broadly at the intersection of AI and the life sciences, we want to connect with the expert scientists, founders, and leaders in the industry. Reach out to Andrew Hedin (ahedin@bvp.com), Marla Jalbut (mjalbut@bvp.com), or Grace Dai (gdai@bvp.com).











